NSF Unidata Science Gateway Update: What's new? What's next?
February 2026
The NSF Unidata Science Gateway is continually evolving to fit the needs of our ever growing community. Since the Spring 2025 semester, 4 new institutions have joined 15 returning ones in taking advantage of the Science Gateway’s custom Python in the Atmosphere and Ocean Science (PYAOS) JupyterHubs for classrooms and research. Our community members challenge us to do new and exciting things, such as launching JupyterHub servers with the capability to run AI/ML workflows, execute WRF and CM1, and launch multi-worker dask clusters.
As the Science Gateway grows in visibility, popularity, and features, we want to share what we’re excited about and some of what’s been going on behind the scenes.
What’s new?
Auto-scaling, automation, and AWIPS, oh my!
The introduction of auto-scaling compute resources to the NSF’s Jetstream2 Cloud has opened many doors for Unidata's science gateway development team (there are two of us). This new feature has prompted some revisions to our JupyterHub provisioning workflow. Our initial investment in automation is already paying off as our efficiency is improved, allowing us to work on more interesting things that benefit you directly! In addition to enabling us to be better stewards of these resources, auto-scaling allows Science Gateway visitors to launch dedicated virtual machines on demand to use established desktop visualization tools such as AWIPS CAVE and NSF Unidata’s Integrated Data Viewer (IDV). We have started equipping every JupyterHub instance with this capability as of Fall 2025. Science Gateway developer Julien Chastang presented this work at the 2026 Conference of the American Meteorological Society. If you missed it and are interested, you can find the slide-deck here.
Sneaky storage improvements
In previous semesters, each JupyterHub user would be allocated their own cloud storage volume, which had to be added to their server on login and removed on logout, much like a physical USB stick. In some frustrating instances, this storage volume would fail to de/attach correctly, leaving users hanging at the login screen. A new shared storage solution has been implemented that circumvents this problem, providing faster JupyterLab start-up times when resources are already available. Most users won’t notice these improvements, but those of you who have been met with the dreaded AttachVolume.Attach failed for volume ... Spawn failed message on login will thank us!
Set sail to new shores with our flagship JupyterHub
One of our goals for 2025 was a relaunch of the Science Gateway’s flagship JupyterHub server found at jupyterhub.unidata.ucar.edu. In the past, this server acted as a demonstration server for interested individuals to visit and check out JupyterHub’s capabilities. However, it was necessary for each visitor to reach out to the Gateway team to be allow listed before logging in. Now, a “federated” identity provider called CILogon lets you log in to the server using your academic institution’s credentials to run notebooks or launch AWIPS CAVE and IDV on-demand. The Science Gateway’s JupyterHub is now an accessible, convenient place to test novel workflows such as those involving NCEPs recently available AIGFS data streams!
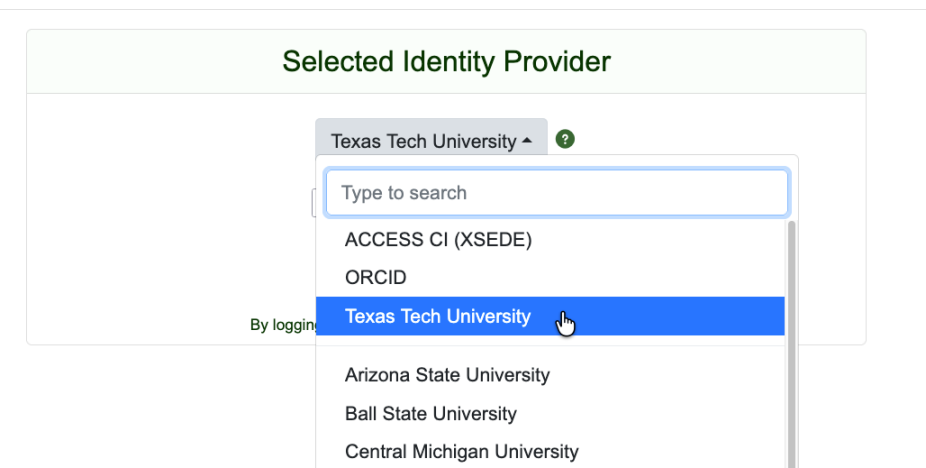
Notes on CILogon
To use the new authentication workflow, browse to our JupyterHub’s login page where you will be prompted to sign in with CILogon. On the CILogon sign in page select your Identity Provider (IdP) and sign in. If your institution is not listed, you have several options:
- Ask us to add your institution to the list of IdPs.
- Ensure your institution is registered with CILogon by searching this page
- Email us at support-gateway@unidata.ucar.edu with your institution’s name
- Create an NSF ACCESS account
- Navigate back to our the login page and select the “ACCESS CI (XSEDE)” IdP
- Contact support (support-gateway@unidata.ucar.edu) to be added to our ORCID or GitHub allow list
If your institution is not registered with CILogon, contact your institution’s IT department and request they register your IdP.
What’s next?
Notebook gallery
One common comment to the NSF Unidata Program Center in our 2025 Community Assessment can be summarized as “I would like some examples or template code of common tasks that I can adapt to my own needs.” NSF Unidata’s next step to address this issue is to leverage the Gateway’s Jupyter notebook capabilities and provide a notebook gallery service. NSF Unidata has some existing example code and notebooks, such as those for MetPy, Siphon, and python-awips. However, these are currently scattered across various documentation sets. For the Gateway's new gallery we envision users being able to not only view and download example notebooks to run on their own compute resources, but to test out example code directly on the Gateway's JupyterHub for rapid interaction and iteration.
Community contributions: Show the world your work
Science and education is about more than access to data and powerful compute environments; there’s a collaborative component to both. A common question that we get from instructors is: “how can I see examples of workflows or classroom exercises from other Gateway users?” Currently the answer is “you can’t!” (🙁) — sharing resources between Gateway users is quite complicated. One of our near-term projects is to move the Gateway beyond isolated instances of the JupyterHub technology and towards a more collaborative space with the addition of a “Community Contributions” feature. We are envisioning this as a space that will allow users from the Gateway’s various JupyterHubs to share, comment on, and access notebooks and computational environments.
We’re hard at work finding the puzzle pieces and putting them together to bring these visions to reality. We look to use technologies from the open source software ecosystem that complement each other, maximizing our efforts and avoiding reinventing the wheel.
BYOE: Bring (and share) Your Own Environments
Given the large diversity of computing packages and libraries required by workflows in the Earth Systems Sciences, one challenge is ensuring that, when authoring or sharing notebooks, users have the necessary packages to run their code. Today, if somebody needs a package, they must reach out to the Gateway team to request it be added to their environment. This is slower than desired and is infeasible for us to do for more users and packages as the Gateway community and feature list grows. We are exploring different methods that will enable Gateway visitors to manage and share their own Jupyter environments. With the click of (hopefully!) a few buttons, users will be able to login to a JupyterHub, fetch a collaborator’s notebook, build a — or access an existing — compute environment, and be off to the races!
Documentation
With a growing repertoire of capabilities, we appreciate the importance of writing and maintaining clear user documentation. As part of a general code-clean-up initiative, the Science Gateway team will be creating a new home for our admittedly sparse documentation, which currently only focuses on the JupyterHub administration aspect of the Gateway.
Closing remarks
All of this is in addition to the less glamorous, more constant, parts of our job: provisioning, tearing down, and maintaining JupyterHub clusters; debugging, troubleshooting, and user support; and testing and improving our DevOps workflows, to name a few. As you can see, we have quite a bit on our plates — don’t we all?. Yet, much like our atmosphere the NSF Unidata Science Gateway is always in motion! The Gateway team is here to serve your needs. If there’s a new feature or improvement you’d like to see implemented we would love to hear from you (support-gateway@unidata.ucar.edu).
Please stay tuned for updates on our progress!
Until next time,
ana victoria espinoza


Add new comment